AbstractPurposePrecision oncology approach for recurrent and metastatic head and neck squamous cell carcinoma (HNSCC) is necessary due to its dismal prognosis. We performed a genomic profile-based umbrella trial of patients with platinum-refractory HNSCC (KCSG-TRIUMPH). Here, we present an in-depth report of the the nintedanib arm (arm 3) of the current trial.
Materials and MethodsThe TRIUMPH study was a multicenter, open-label, single-arm phase 2 trial, in which patients were assigned to treatment arms based on next-generation sequencing (NGS)–based, matching genomic profiles. Patients whose tumors harbor fibroblast growth factor receptor (FGFR) alteration were enrolled in the nintedanib arm (arm 3) as part of the TRIUMPH study. The primary endpoint was the overall response rate (ORR), and secondary endpoints included overall survival (OS), progression-free survival (PFS), safety, and biomarker analysis.
ResultsBetween October 2017 and August 2020, 207 were enrolled in the TRIUMPH study, and eight were enrolled in the nintedanib arm. ORR and disease control rate were 42.9% and 57.1%, respectively. The median PFS was 5.6 months and the median duration of response was 9.1 months. Median OS was 11.1 months. One patient maintained the partial response for 36 months. Overall, the toxicity profiles were manageable.
ConclusionSingle-agent nintedanib has demonstrated significant efficacy in FGFR-mutated, recurrent or metastatic HNSCC patients, with tolerable toxicity profiles. The results from the study have provided the basis for routine NGS screening and FGFR-targeted therapy. Because of the small number of patients due to slow accrual in this study, further studies with a larger cohort are warranted for statistical power.
IntroductionThe current mainstay of treatment for recurrent and/or metastatic head and neck squamous cell carcinoma (HNSCC) involves anti–programmed death-1 immune checkpoint inhibitors with or without platinum-based chemotherapy [1]. Apart from the epidermal growth factor receptor (EGFR) monoclonal antibody, cetuximab, there is currently no established, evidence-based targeted therapy effective in HNSCC [2]. New identification of several molecular targets for HNSCC, with potentially actionable targets including PIK3CA, CDKN2A, EGFR, CCNDA, and fibroblast growth factor receptor (FGFR) has been possible due to advancements in molecular profiling techniques, including next-generation sequencing (NGS), genotyping, and mRNA expression analysis [3], yet studies that validate the role for such targets as predictive biomarkers are hard to find [4].
Thus, a precision oncologic approach in HNSCC to identify patients who are likely to benefit from molecular-targeted therapy was launched as a multicenter, umbrella, phase 2 trial. The TRanslational bIomarker-driven Umbrella Project for Head and neck squamous cell carcinoma (TRIUMPH) study, conducted in association with the Korean Cancer Study Group (KCSG), was the first umbrella trial for platinumre-fractory HNSCC patients, with multiple targeted therapies for phosphoinositide 3-kinase (PI3K), EGFR, FGFR, CDK4/6, and programmed death-ligand 1 (PD-L1)/cytotoxic T-lymphocyte associated protein 4 (CTLA-4) (NCT #03292250 [5]). Here, we report in-depth results for a subset of patients whose tumor harbored alterations (FGFR amplification, mutation, or FGFR3 fusion) in the FGFR pathway, treated with nintedanib in the TRIUMPH trial.
According to The Cancer Genome Atlas (TCGA) data [6], amplifications and mutation of FGFR1, FGFR2, and FGFR3 are reported to occur in 10%, 2%, and 2% of patients with human papillomavirus–negative cancers, respectively. Other studies have reported FGFR genomic alterations, most frequently focal amplifications in FGFR1 and FGFR3 mutations, in approximately 8.9%-37.3% of HNSCC [4,7,8]. Because they are one of the most prevalent receptor tyrosine kinase mutations in HNSCC, the FGFR pathway may serve as a potential, promising therapeutic target [7].
Nintedanib (BIBF1120), an orally available small-molecule tyrosine kinase inhibitor, targets major angiogenesis pathways by inhibiting the FGFR family as well as the platelet-derived growth factor receptor family and vascular endothelial growth factor receptor family kinases [9]. Both in vivo and xenograft experiments have demonstrated good antitumor efficacy, leading to a substantial delay of tumor growth or tumor regression, and histological examination of treated tumors showed a marked reduction of the tumor vessel density up to 80% [10]. Nintedanib has been previously investigated in other solid cancers, including non–small cell lung cancer (NSCLC), as well as for idiopathic pulmonary fibrosis [11-13]. Since 2014, nintedanib has been approved by the European Medicines Agency for use in NSCLC. It is also approved for use in idiopathic pulmonary fibrosis and interstitial lung disease by the United States Food and Drug Administration (US FDA) but not approved for use in any solid cancer [14].
In this phase II, multicenter, single-group clinical trial, as part of a larger umbrella trial, we aim to evaluate the comprehensive efficacy and safety of nintedanib in patients with recurrent or metastatic HNSCC with FGFR pathway alterations, upon failure to respond to platinum-based therapy.
Materials and Methods1. Study design and treatment allocationThe TRIUMPH was a multicenter, investigator-initiated, open-label, phase II umbrella trial consisting of five targeted therapies based on matched biomarkers, for patients with histologically or cytologically confirmed recurrent or metastatic HNSCC [5].
During the screening process, each patient’s mutation profiling in search of driver alterations was assayed via multiplexed targeted NGS assay and Nanostring assay based on tumor RNA expression. Genomic alterations of the PIK3CA, EGFR, FGFR, and cell cycle pathway mutation and amplification were included as the target driver mutations, and patients were assigned to treatment arms per decisions by the molecular tumor board (MTB), dedicated specifically to the TRIUMPH trial.
In brief, the targeted therapies evaluated in this umbrella trial were as follows: arm 1, alpelisib (BYL719), a PI3K inhibitor; arm 2, poziotinib, an EGFR/human epidermal growth factor receptor 2 inhibitor; arm 3, nintedanib, an FGFR inhibitor; arm 4, abemaciclib, a CDK4/6 inhibitor. In case there were no matching targets, the patients were allocated to arm 5, durvalumab. For patients initially enrolled in arm 1-4, crossover to arm 5 upon disease progression was allowed. Tremelimumab was added (arm 5-1) to the durvalumab monotherapy (arm 5) upon disease progression to test if the addition of a CTLA-4 inhibitor could convert non-immunogenic tumors into immunogenic tumors.
2. Molecular profiling assaysNGS assays are described in detail in our previous report of the feasibility study done prior to the TRIUMPH study [15]. Briefly, genomic DNA was isolated from formalin-fixed paraffin-embedded (FFPE) samples using the QIAamp DNA FFPE Tissue Kit (Qiagen, Hilden, Germany) for the targeted sequencing of 244 head and neck cancer-related genes. Library preparation was carried out using customized SureSelectXT Target Enrichement library generation kit (Agilent, Santa Clara, CA), then the libraries were sequenced using the high-throughput, Illumina HiSeq2500 platform with a depth of coverage > 1,000×.
The resulting short reads were quality trimmed using Sickle [16] and mapped to the human reference genome using Burrows-Wheeler Aligner [17], and the aligned reads were then processed with Genome Analysis ToolKit v3.5 [18]. Variant detection was carried out using MuTect v1.17 and Varscan2 v2.3.7 and ANNOVAR was used for functional annotation of the high confidence variants. Copy number alterations were called using CNVkit and genes with > 4 and 0 measured copy numbers were considered amplified and deleted, respectively.
Additionally, Nanostring assay for RNA expression analysis was performed by first isolating total tumor RNA with the RNeasy kit (Qiagen). Using nCounter Analysis System (Nanostring Technologies, Seattle, WA), the expression of 55 immune-related genes was screened.
3. Patient selectionPatients with genetic alterations in the FGFR pathway were eligible to enroll in the nintedanib arm of the trial. Eligible patients were those with histologically confirmed HNSCC; recurrent and/or metastatic HNSCC of the oral cavity, oropharynx, hypopharynx, larynx, nasal cavity, maxillary sinus; not amenable to curative treatment and had progressed or recurred upon one or two systemic therapy regimens including platinum-based chemotherapy, or within 6 months after concurrent chemo-radiation (CCRT) administered as definitive treatment, with or without induction chemotherapy; aged ≥ 20 years; at least one measurable disease according to Response Evaluation Criteria in Solid Tumor (RECIST) ver. 1.1; an Eastern Cooperative Oncology Group performance status of 0 or 1; and adequate organ function. Key exclusion criteria included patients previously treated with FGFR pathway inhibitors; nasopharyngeal carcinoma; active brain metastasis; previous surgery requiring general anesthesia within 4 weeks before enrollment, and history of ileus, airway obstruction, or active bleeding within 6 months of enrollment.
Patients in the FGFR arm were treated with a single-agent nintedanib at a starting dose of 200 mg twice daily, and upon progression were allowed crossover to durvalumab monotherapy arm (arm 5) of the TRIUMPH trial.
4. Treatment and assessmentPatients with FGFR mutation/amplification uncovered by the tissue (archival or fresh tumor tissue)-based NGS were enrolled and received 200 mg of nintedanib twice daily through oral administration in 28-day cycles until disease progression, unacceptable toxicity, or withdrawal of consent. Dose modifications or delays were considered based on the worst grade of toxicity, according to the approved protocol.
Tumor response was evaluated according to the RECIST ver. 1.1 at baseline and every 8 weeks thereafter. Treatment beyond progression was not allowed, as crossover was permitted. If patients crossed over to the durvalumab arm upon progression, 750 mg of durvalumab was administered intravenously every 2 weeks, up to a total cycle of 18 cycles. Safety was evaluated in patients who received at least one dose of treatment, and adverse events (AEs) were evaluated using the National Cancer Institute Common Terminology Criteria for Adverse Events ver. 4.03.
Patients at every visit at the start of each treatment cycle underwent laboratory tests for complete blood count, plasma biomarkers, and serum metabolic panel.
5. Statistical analysisThe primary endpoint was the investigator-assessed objective response rate (ORR), that is, the percentage of patients who achieved an objective response of either complete response (CR) or partial response (PR) according to RECIST ver. 1.1. Secondary endpoints included overall survival (time from first treatment to death), progression-free survival (time from first treatment to disease progression, death, or last tumor assessment, whichever comes first), safety, and biomarker analysis. The sample size was calculated using the one-arm binomial method, with the null hypothesis that the ORR was ≤ 5% against the alternative hypothesis that the ORR was ≥ 20%, at a significance level of 10%. The hypothesis was tested with 90% statistical power and a 10% significance level, and a total of 29 patients are required for response assessment. Considering dropout rates of 10%, the final sample size was calculated to include 32 patients.
The efficacy set was defined as patients who had baseline tumor assessment and had received at least one dose of study treatment. The Clopper-Pearson estimation method was used to calculate 95% confidence intervals (CIs) for the proportion of patients with an objective response. Median progression-free survival, overall survival, and their 95% CIs were estimated using the Kaplan-Meier method in the efficacy analysis population.
Safety analyses were performed on the safety set of all treated patients, and AEs were summarized based on the frequency and proportion of patients according to the preferred term. Statistical analyses were completed using SAS ver. 9.4 (SAS Institute Inc., Cary, NC) or R ver. 3.4.4 (R Foundation for Statistical Computing, Vienna, Austria).
Results1. Patients characteristicsBetween October 2017 and August 2020, a total of 180 HNSCC patients were screened for the TRIUMPH study, and ultimately 207 patients were enrolled, including those enrolled in the crossover arm (arm 5, durvalumab) (Fig. 1A). Based on the results of the MTB, 10 patients were initially assigned to the nintedanib arm (arm 3) and eight patients were ultimately enrolled (Fig. 1B). Two patients initially enrolled in the nintedanib arm later crossed over to receive durvalumab upon progression.
At the time of data cutoff on December 2021, all had discontinued treatment—three had completed the study, four had died, and one had withdrawn from the study upon patient request. The median follow-up period from the start of treatment was 29.5 months. Due to the lower-than-expected rate of FGFR alterations resulting in poor accrual, patient enrollment in the nintedanib arm of the TRIUMPH study was terminated prematurely.
Patient baseline characteristics are described in Table 1. The primary site of tumor included those in the oropharynx (12.5%), as well as the hypopharynx (25%), larynx (12.5%), and nasal cavity (37.5%), and the majority of patients had received prior surgery or radiotherapy and had received up to three previous lines of systemic therapy. All patients enrolled in this arm met the eligibility criteria and were included in the safety analysis; seven of the eight were included in the efficacy analysis.
2. ResponseThree out of seven patients available for efficacy analysis achieved PR, bringing the overall response rate to 42.9% (95% CI, 9.9 to 81.6). The disease control rate was 57.1% (95% CI, 18.4 to 90.1), which included one patient with stable disease for 11.0 months. Efficacy results are summarized in Table 2. The median progression-free survival (PFS) was 5.6 months (95% CI, 0.92 to 14.6 months) (Fig. 2A), and the median duration of response (DoR) was 9.1 months (95% CI, 3.75 to not reached). Median overall survival (OS) was 11.1 months (95% CI, 1.9 to not reached) (Fig. 2B). Fig. 3A and B depicts the waterfall plot showing the magnitude of the best tumor response from baseline and swimmer’s plot in patients who were available for efficacy analysis.
Here we report three cases of HNSCC patients achieving PR on nintedanib treatment.
(1) Case 1The patient with the longest DoR is a 38-year-old female with nasal cavity cancer who had previously undergone endoscopic medial maxillectomy and had progressed upon two previous lines of systemic therapy comprised of DCF (docetaxel, 5-fluorouracil, cisplatin) regimen and immunotherapy (CD47, pembrolizumab). Upon whole-exome and Nanostring assay, her tumor was found to harbor FGFR1 frameshift deletion (p.W190fs, p.W182fs, p.W188fs, p.W221fs, p.W101fs, and p.W99fs).
The patient received nintedanib from August 2018 to August 2021 and achieved PR (tumor shrinkage of 81.4% from baseline) according to radiologic response assessment by RECIST ver. 1.1. Although she has been reported as a partial responder with only a minimal trace of disease remaining, clinically she is deemed to be in CR. Interestingly, the patient has been on close observation after completion of nintedanib treatment and has been progression-free at the time of manuscript writing. Gross tumor morphology and radiologic disease response are depicted in Fig. 4.
(2) Case 2This is a 57-year-old male with poorly differentiated hypopharyngeal carcinoma, with initial stage IV diagnosis and FGFR1 amplification (CNV 8). After initial CCRT and a prior line of systemic chemotherapy, he was enrolled in the nintedanib arm of the TRIUMPH trial. Although notable tumor shrinkage was achieved (46.4% reduction from baseline), ultimately disease progressed after 5.6 months, and the patient passed away 6 months later.
(3) Case 3This is a 72-year-old male with hypopharyngeal carcinoma who, upon NGS, was found to harbor various mutations—TP53, KDM5C, PTCH1, KMT2C, DDR2, KDM5A, and FBN2, to name a few, as well as FGFR3 with multiple single nucleotide variation sites (Fig. 5B). The patient received nintedanib therapy and achieved tumor shrinkage (46.2% reduction from baseline tumor size according to RECIST ver. 1.1), with a durable response for more than a year (14.6 months) before ultimate disease progression, with the OS of 23 months.
3. SafetySafety analysis was performed in all nintedanib-treated patients (n=8). Treatment-emergent adverse events arose in seven out of eight patients (87.5%), and a serious adverse event was noted in one patient (12.5%). Adverse drug reactions were reported in five patients (62.6%), yet no serious adverse drug reaction occurred.
The most common AEs regardless of grade were diarrhea (4/8, 50.0%), followed by anorexia (3/8, 37.5%) and increased alanine aminotransferase levels (2/8, 25.0%). Grade 3 AEs occurred in three different patients—which included increased aspartate aminotransferase levels, diarrhea, and noncardiac chest pain (12.5%, 12.5%, and 12.5%, respectively). Table 3 outlines the AEs. There was no new significant safety signal beyond the previous clinical trials with nintedanib [13,19].
Two patients (25.0%) underwent dose delay and ultimately dose reductions because of the AEs, but one event was due to a bone fracture deemed unlikely to be caused by the drug. Most AEs were manageable with adequate treatment, and no study discontinuation due to AEs of treatment-related deaths had occurred.
4. Mutation profiling of the FGFR familyHere, we present a further analysis of the mutation profiles of the patients initially screened and assigned to the nintedanib arm (n=10). Per protocol, patients harboring FGFR amplifications or mutations were eligible for enrollment. Six patients (6/10, 60%) were found to have FGFR1 mutations/amplifications, five of which were amplifications. One patient harboring FGFR1 frameshift deletions, as introduced in the prior section (case 1), showed exceptional response to nintedanib.
Other mutations in the FGFR pathway included FGFR2 (1 amplification, 10%), FGFR3 (1 amplification and one multihit mutation at p.G380R, p.G382R, p.D581N, p.D580N, p.D468N, p.D582N, and p.E685X[stopgain]), and FGFR4 overexpression revealed by the Nanostring assay for RNA expression analysis in one patient. Detailed mutation profiling of the nintedanib arm sub-cohort and a diagram showing the distribution of different FGFR3 mutation sites [20] is shown in Fig. 5.
Concurrent oncogenic mutations found with FGFR mutations included genes related to the PIK3CA pathway (PIK3CA amplification), EGFR pathway (multiple mutations at p.D792N, p.D784N, p.D570N, and p.D837N), as well as the cell cycle pathway (CCND1 and CCND2 amplification, and CDKN2A deletion).
DiscussionHere we report the results of oral multi-kinase inhibitor nintedanib on a biomarker-driven subset cohort, as part of a larger precision-oncology, multi-arm umbrella trial (TRIUMPH) in HNSCC. While the TRIUMPH umbrella study in general did not meet the pre-specified primary endpoint, it has proven valuable points in that next-generation sequencing-based phenotyping is a valid option for matching targeted agents in recurrent or metastatic HNSCC.
Identification and analyzing exceptional responders, as defined by the National Cancer Institute (NCI), could provide a valuable asset as a basis for further molecular studies [21]. Among the three cases presented above, the first patient exhibited an exceptional response to nintedanib. Despite whole-exome and Nanostring-based RNA analysis, the precise mechanism underlying the exceptional response remains unclear. Further investigations are warranted to elucidate the underlying biological factors that contributed to the patient’s remarkable response to nintedanib therapy.
In the nintedanib-treated patients enrolled in the current trial, the primary endpoint of ORR was met, with 42.9% against the initial hypothesis of 20%, with disease control rate also reaching 57.1% (4/7). However, this nintedanib sub-cohort holds some limitations. We initially conjectured that the prevalence of FGFR alteration in our large-scale, NGS-based sequencing of the HNSCC cohort would be comparable to the FGFR mutation rate reported by the TCGA group, in which FGFR1, FGFR2, and FGFR3 mutations have been reported to occur in 10%, 2%, and 2%, respectively [6]. Due to a lower-than-expected number of patients with FGFR mutations and the ensuing slow patient accrual, regrettably the study had to be terminated prematurely.
Previously, nintedanib has been tested in various solid tumors that exhibit a notable incidence of FGFR alterations [22], such as urothelial cancer (31.7%) [23], ovarian cancer (8.6%) [24,25], and NSCLC (5.2%) [11,13], regardless of the presence or absence of FGFR target mutations. One such study was done on salivary gland tumors in a total of 20 patients, and while nintedanib did not yield a PR, the study achieved a 75% disease control rate (15 out of 20 patients) [26]. Other studies have explored the efficacy of nintedanib in combination with other chemotherapy agents in various settings. In phase III, LUME-Lung 1 trial, nintedanib and docetaxel combination showed improved PFS (3.4 vs. 2.7 months; hazard ratio [HR], 0.79 [95% CI, 0.68 to 0.92]) and OS (10.9 vs. 7.9 months; HR, 0.75 [95% CI, 0.60 to 0.92]) compared with docetaxel alone as second-line therapy in NSCLC patients [11]. In a similar study evaluating the efficacy of nintedanib and pemetrexed combination as second-line treatment in NSCLC (LUME-Lung 2 trial [13]), combination treatment reportedly showed significantly prolonged PFS over pemetrexed alone (median, 4.4 months vs. 3.5 months; HR, 0.83 [95% CI, 0.70 to 0.99]; p=0.0435).
What these previous studies have yet failed to answer is whether biomarker-driven patient selection improves FGFR inhibitor sensitivity. There is some evidence from translational research using FGFR2-amplified gastric cancers that point toward FGFR amplification playing a key role in response to FGFR inhibitors. Only high copy-number (high-level) gene amplification translates into high mRNA and protein expression which results in PI3K and mammalian target of rapamycin (mTOR) signaling pathway becoming dependent on FGFR signaling, thus responding to selective FGFR inhibition [27]. While in overall, significant response rate was achieved by selecting patients based on FGFR mutation profile, neither the degree of FGFR amplification determined by copy number variation nor FGFR mRNA overexpression showed clearcut correlation with nintedanib response in our study.
Taking a deeper, comprehensive look at the responders in our trial, one interesting point to note is that none of the responders had concurrent mutation sites in either the EGFR, PIK3CA or the cell cycle pathway. We can further conjecture that having these concurrent mutations may somehow play a role in bypassing the antitumor activities of nintedanib and in the resistant mechanism, but further studies are needed to verify such a hypothesis.
So far, there are several FGFR inhibitors currently approved by the US FDA, such as erdafitinib (JNJ-42756493, Balversa) in bladder cancer [28], pemigatinib (INCB054828, Pemazyre) [29], infigratinib (BGJ398, Truseltiq) [30], and futibatinib (TAS-120, Lytgobi) [31] for use in biliary tract cancer. Others are still in the process of ongoing clinical trials, but the majority are tissue-agnostic trials on hematologic/solid cancers harboring FGFR gene aberrations [32]. To our knowledge, only a handful of biomarker-driven FGFR inhibitor trials are focused solely on HNSCC (NCT04203719, NCT03088059).
As the first biomarker-driven umbrella trial for HNSCC patients who have progressed on previous platinum-based therapy, our study provides further insight into the clinical utility of an NGS-based, precision oncology approach to targeted treatment. This study may also pave way for further research into targeting the FGFR pathway, and better-tailored patient selection for enhanced responses.
In conclusion, single-agent nintedanib showed promising efficacy, with durable response and disease stabilization in FGFR-altered, recurrent or metastatic HNSCC patients, with tolerable toxicity profiles. The results from the study have proved the utility of NGS screening in HNSCC patients. However, because of limited patient enrollment for this study, further prospective studies for nintedanib with an expanded number of patients are highly warranted for more cumulative evidence on nintedanib.
NotesEthical Statement The TRIUMPH study was conducted following the Declaration of Helsinki and the International Conference on Harmonization Guidelines for Good Clinical Practice. The institutional review board of each trial center approved the protocol. Patients provided written informed consent before participation and the index patient included in the manuscript agreed on providing the photography. This study was registered at ClinicalTrials.gov (NCT03292250). Author Contributions Conceived and designed the analysis: Lim SM, Keam B, Kim JS. Collected the data: Kim KH, Ahn HK, Lee YG, Lee KW, Ahn MJ, Keam B, Kim HR, Lee HW, An HJ, Kim JS. Contributed data or analysis tools: Lim SM, Ahn HK, Lee YG, Lee KW, Ahn MJ, Keam B, Kim HR, Lee HW, An HJ, Kim JS. Performed the analysis: Kim KH, Keam B, Kim HR, Kim JS. Wrote the paper: Kim KH, Lim SM, Kim JS. Conflicts of Interest This study was supported by Boehringer Ingelheim (BI). BI had no role in the design, analysis or interpretation of the results in this study. BI was given the opportunity to review the manuscript for medical and scientific accuracy as it relates to BI substances, as well as intellectual property considerations. The authors declare that there is no conflict of interest. AcknowledgmentsThe TRIUMPH study was supported by a grant from the National R&D Program for Cancer Control, Ministry of Health and Welfare, Republic of Korea (grant number HA16C0015). The funder had no role in the study design, data collection, analyses, and interpretation of data, in the writing of the report, or in the decision to submit the article for publication.
Fig. 1.Study scheme and consort diagram for nintedanib arm. (A) Overall scheme of the main TRIUMPH study. (B) Consort diagram showing trial profile for nintedanib arm (arm 3). EGFR, epidermal growth factor receptor; FGFR, fibroblast growth factor receptor; HER2, human epidermal growth factor receptor 2; HNSCC, head and neck squamous cell carcinoma; PD-L1, programmed death-ligand 1; PI3K, phosphoinositide 3-kinase. 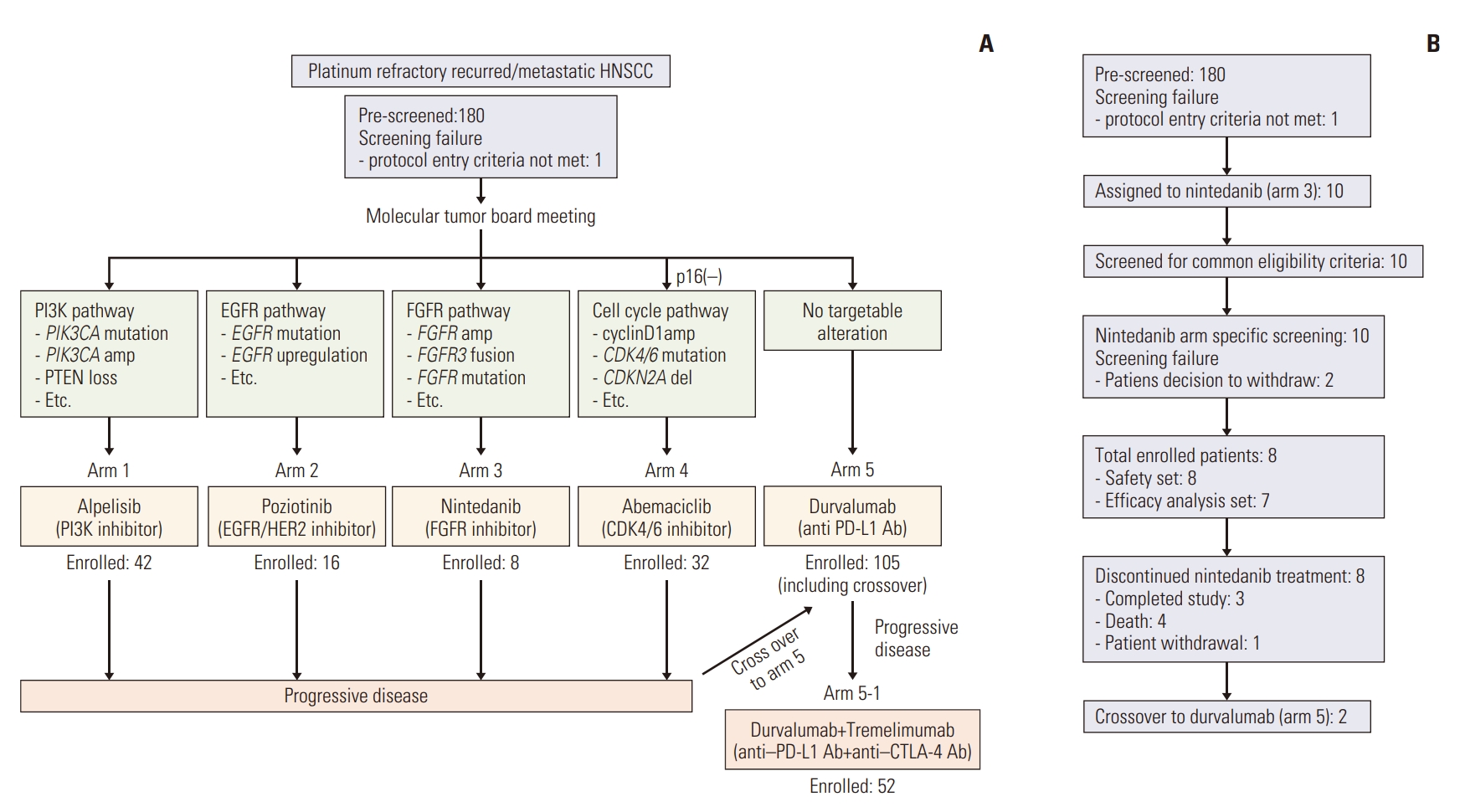
Fig. 2.Kaplan-Meier survival curves. Progression-free survival (A) and overall survival (B) in patients with fibroblast growth factor receptor (FGFR) mutation/amplification treated with nintedanib in the efficacy set (n=7). CI, confidence interval. 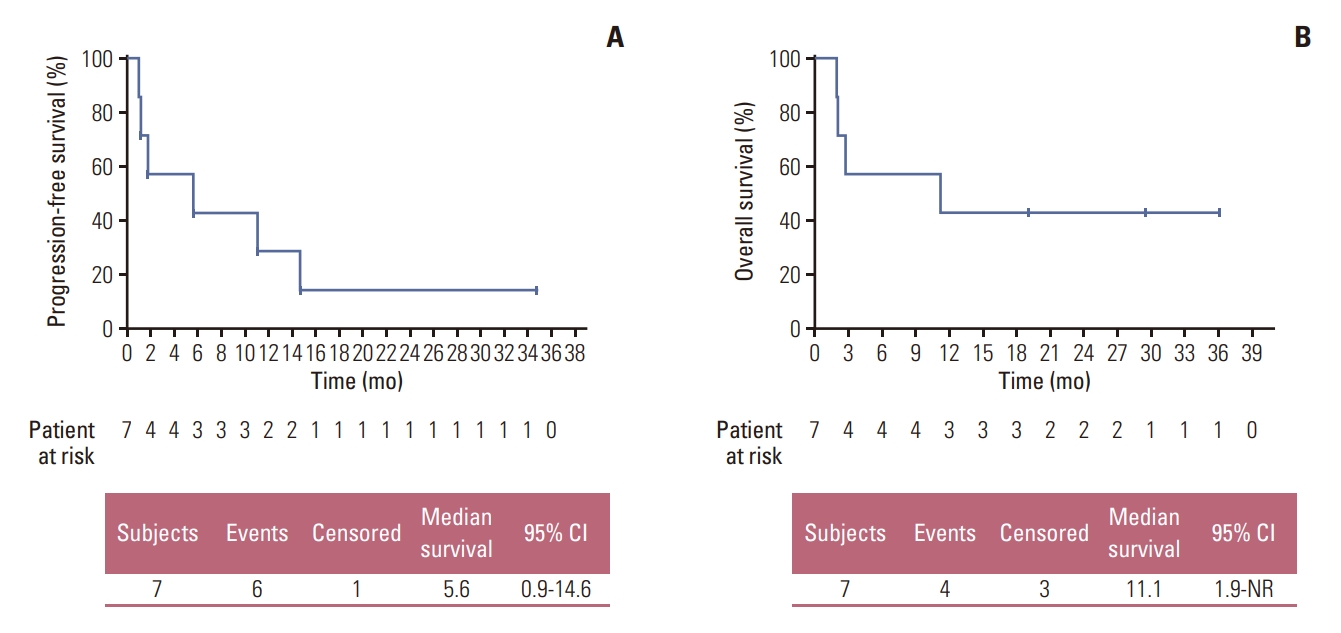
Fig. 3.Waterfall and swimmer plot showing patient tumor response. (A) Waterfall plot for best percentage changes from baseline. (B) Swimmer plot for time on treatment by patient. FGFR, fibroblast growth factor receptor. 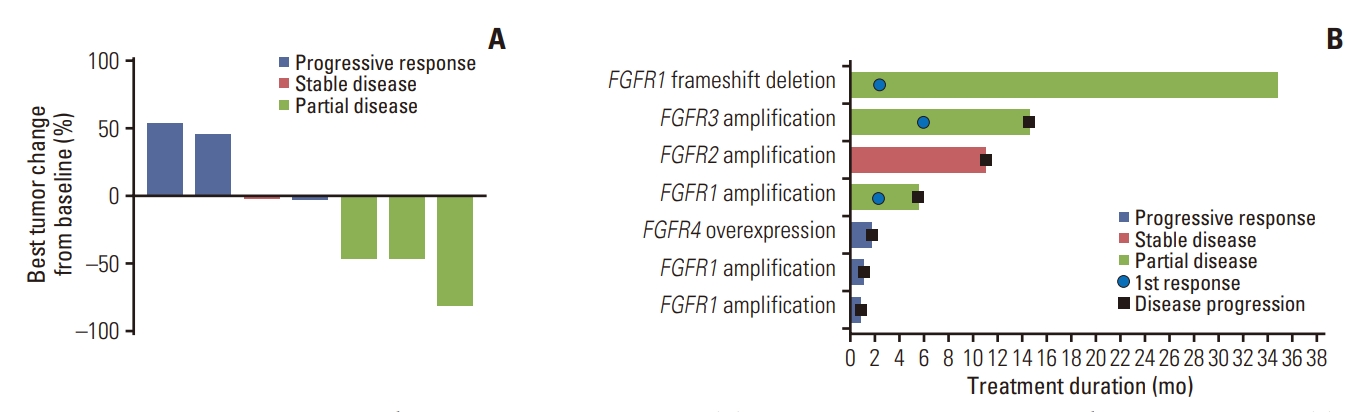
Fig. 4.Nintedanib response in a nasal cavity cancer patient. (A) Gross protruding nasal cavity mass at baseline, after 1st cycle (4 weeks since start of therapy), and at 1.5 years from treatment completion (4.5 years since start of therapy). (B) Neck computed tomography scans at baseline, at 1st response assessment (at 8 weeks), and after 2.5 years of treatment. 
Fig. 5.Integrated clinical and genomic profile of patients screened for the nintedanib arm. (A) Clinical response and Oncoplot showing both patient-specific mutations in fibroblast growth factor receptor (FGFR) pathway as well as other, concurrent mutations in the major tumor-related molecular pathways. Copy number variations (CNV) are annotated. for each patient with FGFR gene amplifications. Nintedanib best response are also given for each patient, with the exception of those who ultimately failed to enroll or dropped out of trial before response evaluation, in which best response are annotated as NA (not available). (B) Lollipop plot showing immunoglobulin-like domains (IG, I-set), tyrosine kinase domain (Pkinase Tyr), and distribution of different mutation sites in single patient with FGFR3 multihit mutations. HPV, human papillomavirus; PD, progressive disease; PR, partial response; SD, stable disease. 
Table 1.Patient baseline characteristics Table 2.Efficacy results in the efficacy set Table 3.Adverse events References2. Vermorken JB, Trigo J, Hitt R, Koralewski P, Diaz-Rubio E, Rolland F, et al. Open-label, uncontrolled, multicenter phase II study to evaluate the efficacy and toxicity of cetuximab as a single agent in patients with recurrent and/or metastatic squamous cell carcinoma of the head and neck who failed to respond to platinum-based therapy. J Clin Oncol. 2007;25:2171–7.
3. Morris LG, Chandramohan R, West L, Zehir A, Chakravarty D, Pfister DG, et al. The molecular landscape of recurrent and metastatic head and neck cancers: insights from a precision oncology sequencing platform. JAMA Oncol. 2017;3:244–55.
4. Marret G, Bieche I, Dupain C, Borcoman E, du Rusquec P, Ricci F, et al. Genomic alterations in head and neck squamous cell carcinoma: level of evidence according to ESMO Scale for Clinical Actionability of Molecular Targets (ESCAT). JCO Precis Oncol. 2021;5:215–26.
5. Keam B, Hong MH, Shin SH, Heo SG, Kim JE, Ahn HK, et al. Personalized biomarker-based umbrella trial for patients with recurrent or metastatic head and neck squamous cell carcinoma: KCSG HN 15-16 TRIUMPH trial. J Clin Oncol. 2023;Sep. 12[Epub]. https://doi.org/10.1200/JCO.22.02786
6. Cancer Genome Atlas Network. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature. 2015;517:576–82.
7. Wang Z, Anderson KS. Therapeutic targeting of FGFR signaling in head and neck cancer. Cancer J. 2022;28:354–62.
8. Seiwert TY, Zuo Z, Keck MK, Khattri A, Pedamallu CS, Stricker T, et al. Integrative and comparative genomic analysis of HPV-positive and HPV-negative head and neck squamous cell carcinomas. Clin Cancer Res. 2015;21:632–41.
9. Roth GJ, Binder R, Colbatzky F, Dallinger C, Schlenker-Herceg R, Hilberg F, et al. Nintedanib: from discovery to the clinic. J Med Chem. 2015;58:1053–63.
10. Hilberg F, Roth GJ, Krssak M, Kautschitsch S, Sommergruber W, Tontsch-Grunt U, et al. BIBF 1120: triple angiokinase inhibitor with sustained receptor blockade and good antitumor efficacy. Cancer Res. 2008;68:4774–82.
11. Reck M, Kaiser R, Mellemgaard A, Douillard JY, Orlov S, Krzakowski M, et al. Docetaxel plus nintedanib versus docetaxel plus placebo in patients with previously treated nonsmall-cell lung cancer (LUME-Lung 1): a phase 3, double-blind, randomised controlled trial. Lancet Oncol. 2014;15:143–55.
12. Richeldi L, du Bois RM, Raghu G, Azuma A, Brown KK, Costabel U, et al. Efficacy and safety of nintedanib in idiopathic pulmonary fibrosis. N Engl J Med. 2014;370:2071–82.
13. Hanna NH, Kaiser R, Sullivan RN, Aren OR, Ahn MJ, Tiangco B, et al. Nintedanib plus pemetrexed versus placebo plus pemetrexed in patients with relapsed or refractory, advanced non-small cell lung cancer (LUME-Lung 2): a randomized, double-blind, phase III trial. Lung Cancer. 2016;102:65–73.
15. Lim SM, Cho SH, Hwang IG, Choi JW, Chang H, Ahn MJ, et al. Investigating the feasibility of targeted next-generation sequencing to guide the treatment of head and neck squamous cell carcinoma. Cancer Res Treat. 2019;51:300–12.
16. Joshi NA, Fass JN. Sickle: a sliding-window, adaptive, quality-based trimming tool for FastQ files (version 1.33) [Internet]. San Francisco, CA: GitHub Inc; 2011. [cited 2023 Mar 1]. Available from: https://github.com/najoshi/sickle
17. Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–60.
18. McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20:1297–303.
19. Otsubo K, Kishimoto J, Ando M, Kenmotsu H, Minegishi Y, Horinouchi H, et al. Nintedanib plus chemotherapy for nonsmall cell lung cancer with idiopathic pulmonary fibrosis: a randomised phase 3 trial. Eur Respir J. 2022;60:2200380.
20. Ou J, Zhu LJ. trackViewer: a Bioconductor package for interactive and integrative visualization of multi-omics data. Nat Methods. 2019;16:453–4.
21. Conley BA, Staudt L, Takebe N, Wheeler DA, Wang L, Cardenas MF, et al. The exceptional responders initiative: feasibility of a National Cancer Institute pilot study. J Natl Cancer Inst. 2021;113:27–37.
22. Helsten T, Elkin S, Arthur E, Tomson BN, Carter J, Kurzrock R. The FGFR landscape in cancer: analysis of 4,853 tumors by next-generation sequencing. Clin Cancer Res. 2016;22:259–67.
23. Hussain SA, Lester JF, Jackson R, Gornall M, Qureshi M, Elliott A, et al. Addition of nintedanib or placebo to neoadjuvant gemcitabine and cisplatin in locally advanced muscle-invasive bladder cancer (NEOBLADE): a double-blind, randomised, phase 2 trial. Lancet Oncol. 2022;23:650–8.
24. Barra F, Lagana AS, Ghezzi F, Casarin J, Ferrero S. Nintedanib for advanced epithelial ovarian cancer: a change of perspective? Summary of evidence from a systematic review. Gynecol Obstet Invest. 2019;84:107–17.
25. Ray-Coquard I, Cibula D, Mirza MR, Reuss A, Ricci C, Colombo N, et al. Final results from GCIG/ENGOT/AGO-OVAR 12, a randomised placebo-controlled phase III trial of nintedanib combined with chemotherapy for newly diagnosed advanced ovarian cancer. Int J Cancer. 2020;146:439–48.
26. Kim Y, Lee SJ, Lee JY, Lee SH, Sun JM, Park K, et al. Clinical trial of nintedanib in patients with recurrent or metastatic salivary gland cancer of the head and neck: a multicenter phase 2 study (Korean Cancer Study Group HN14-01). Cancer. 2017;123:1958–64.
27. Pearson A, Smyth E, Babina IS, Herrera-Abreu MT, Tarazona N, Peckitt C, et al. High-level clonal FGFR amplification and response to FGFR inhibition in a translational clinical trial. Cancer Discov. 2016;6:838–51.
28. Loriot Y, Necchi A, Park SH, Garcia-Donas J, Huddart R, Burgess E, et al. Erdafitinib in locally advanced or metastatic urothelial carcinoma. N Engl J Med. 2019;381:338–48.
29. Abou-Alfa GK, Sahai V, Hollebecque A, Vaccaro G, Melisi D, Al-Rajabi R, et al. Pemigatinib for previously treated, locally advanced or metastatic cholangiocarcinoma: a multicentre, open-label, phase 2 study. Lancet Oncol. 2020;21:671–84.
30. Javle M, Roychowdhury S, Kelley RK, Sadeghi S, Macarulla T, Weiss KH, et al. Infigratinib (BGJ398) in previously treated patients with advanced or metastatic cholangiocarcinoma with FGFR2 fusions or rearrangements: mature results from a multicentre, open-label, single-arm, phase 2 study. Lancet Gastroenterol Hepatol. 2021;6:803–15.
|
|
|||||||||||||||||||||||||||||||||||||||||